Distinguished Professor Sue Lunte named December 2024 Sutton Family Research Impact Award recipient

Above: Sue Lunte and Emily Kurfman
The Department of Chemistry congratulates Distinguished Professor Sue Lunte on receiving the December 2024 Sutton Family Research Impact Award!
The Sutton Award is a monthly competition among chemistry faculty. Every month, the Chemistry Department Chair and Associate Chairs review the peer-reviewed papers published by chemistry faculty from the three previous months to select a winner. The recipient receives a $500 cash prize and is featured on the departmental website.
For a full list of winners, visit our Sutton Family Research Impact Award webpage.
Development of capillary electrophoresis methods for detection of microbial metabolites on potential future spaceflight missions
By Emily A. Kurfman, Maria F. Mora, Peter A. Willis, and Susan M. Lunte
Electrophoresis 2024; 45:1684-1691. DOI: 10.1002/elps.202400073
Future space exploration missions will likely focus on ocean worlds, such as Europa (a moon of Jupiter) and Enceladus (a moon of Saturn), to search for possible signatures of life associated with their liquid water oceans. To detect and profile the chemical composition of single-celled life that might be discovered, it is necessary to develop methods that are spaceflight compatible for in situ analysis. In particular, methods for the separation and detection of small organic molecules that could be signs of life are needed. Capillary electrophoresis (CE) is a useful separation method for determination of charged analytes of biological significance, including amino acids and nucleobases. An instrument based on CE in combination with multiple detection systems, including laser induced fluorescence (LIF) and mass spectrometry (MS), is currently under development for spaceflight applications. In this paper, we report CE-based methods for separation and detection of major polar metabolites in cells, such as amino acids, nucleobases/sides, and oxidized and reduced glutathione using detectors that are less expensive alternatives to LIF and MS. Direct UV detection, indirect UV detection, and C4D were tested, and a combination of direct UV and C4D allowed the detection of the widest variety of metabolites. The optimized method was used to profile metabolites found in samples of E. coli and P. haloplanktis and showed distinct differences between the species.

Fig. 1: Europa
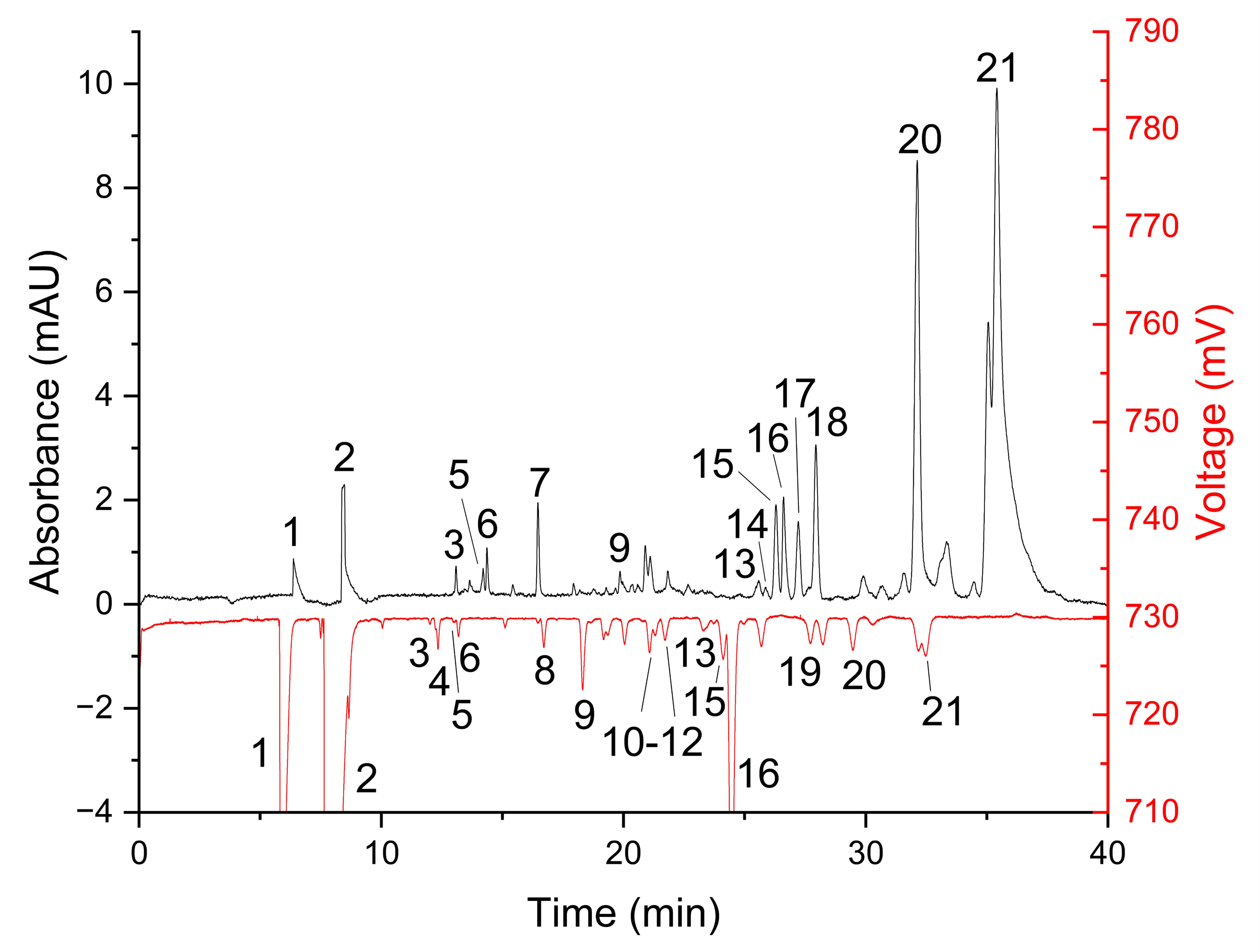
Fig. 2: Capillary electrophoresis (CE)-UV at 200 nm (top trace) and CE-C4D (bottom trace) analysis of an extract in 80% (v/v) methanol of a Pseudoalteromonas haloplanktis culture sample. Likely peak identities: 1 = K+, 2 = Na+ and/or Mg2+, 3 = cytosine, 4 = Lys, 5 = adenine and/or Arg, 6 = His, 7 = guanine, 8 = Gly, 9 = adenosine + Ala, 10 = Val, 11 = Ile, 12 = Leu and/or Ser, 13 = Asn, 14 = Met, 15 = Trp and/or Gln, 16 = Glu, 17 = Phe, 18 = Tyr and/or Pro, 19 = Asp, 20 = GSSG, and 21 = GSH. Conditions: background electrolyte (BGE) = 2 M acetic acid, field strength = 235 V/cm, capillary effective length = 55 cm at C4D detector and 60.75 cm at UV detector, and UV detection wavelength = 200 nm.